

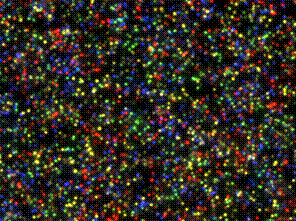

What is it? : Polony = Polymerase-colony (pronounced "Pahl'ahnee" at Hahvahd, and "Pole own' ee" at the Deli).
What flavors? (1) in situ slide PCR, (2) emulsion-PCR-beads, (3) Isothermal amplification (no-PCR, no-gels, no-beads).
How to get it? (open source)
Software: 2009.
(Image analysis for in-situ haplotyping, exon-typing 2003 , Polony-bead software 2005,
2006).
Hardware: 2009: Danaher Polonator G.007 (2003 & 2006 : Slide amplifiers, fuidics, scanners or bead microscope 2005).
Wetware: Protocols 2008 OWW.
(2003. Cleavable Fluor-dNTPs)
Shareware: Public discussion forum Polony-Protocols
ESL-ware: Ethical, Legal, Social innovation in shareable medical research & frank/detailed discussion of re-identification issues:
Personal Genome Project.
Tuneware: mp3 : James Kugler
Brought to you by: Advanced Sequencing Technology NHGRI grants
CEGS ,
2004 & 2005
Ancient History of:
Nucleic Acid Cloning and Sequencing by Synthesis.
Costs: Polony cost details
; Cost-Error Comparison Method
; Comparison spreadsheet
; 454 vs ABI-3730xl
Disclosure: Church lab licenses & advisory roles to 14 of the 17 companies below.
Multiplex Cyclic Sequencing by Synthesis References
A. Illumina/Solexa
3. Porreca GJ, Zhang K, Li JB, Xie B, Austin D, Vassallo SL, LeProust EM, Peck BJ, Emig CJ, Dahl F, Yuan Gao Y, Church GM, Shendure, J (2007) Multiplex amplification of large sets of human exons. Nat Methods. 2007 Nov;4(11):931-6.
2. Johnson DS, Mortazavi A, Myers RM, Wold B. (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science 316(5830):1441-2.
1. Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K.(2007) High-resolution profiling of histone methylations in the human genome. Cell. 129(4):823-37.
B. ABI/APG SOLiD
4. http://bioinformatics.oxfordjournals.org/cgi/content/full/24/23/2776
3. http://genome.cshlp.org/content/early/2009/06/18/gr.091868.109.abstract
2. http://www.ncbi.nlm.nih.gov/pubmed/18775913
1. http://www.ncbi.nlm.nih.gov/pubmed/18516046
C. 454-Roche
2.-etc. Other publications
1. Margulies, M. Eghold, M. et al. (2005) Genome sequencing in microfabricated high-density picolitre reactors Nature. Sep 15; 437(7057):326-7
D. Complete Genomics >95% Human genome sequencing service -- instruments not for sale.
E. IntelligentBioSystems (IBS)
F. Helicos Biosciences Corp -- Single molecule Polymerase SbP
G. Pacificbiosciences -- Single molecule Polymerase SbP
H. Visigen -- Single molecule Polymerase
I. LightSpeed Genomics - High resolution SbL
J. Bionanomatrix -- Fluorescent SbP
K. Genizon Biosciences Cantaloupe - Fluorescent SbH
L. Ion Torrent Systems - Chemical Field-Effect Transitor detection of Polymerase dNTP cleavage
M. Danaher/Dover/Azco Polonator G.007 Contact (related references below)
21. Forest CR, Rosenbaum AM, Church GM (2008) DNA Sequencing By Ligation On Surface-Bound Beads In A Microchannel Environment MicroTAS 2008
20. Church GM, Porreca GJ, Terry RC, Lares M (2008) High-speed imaging for DNA sequencing. Biophotonics.
19. Nardi V, Raz T, Chao X, Wu CJ, Stone RM, Cortes J, Deininger MWN, Church G, Zhu J and Daley GQ (2008) Monitoring resistance to kinase inhibitors with polony technology: towards personalized therapy for CML patients. Oncogene 27(6):775-82.
18. Conrad C, Jun Zhu J, Conrad C, Schoenfeld D, Fang Z, Ingelsson M, Stamm S, Church G, Hyman BT (2007) Single Molecule Profiling of tau Gene Expression in Alzheimer's Disease. J. Neurochem 103(3):1228-36.
17. Kim JB, Porreca GJ, Gorham JM, Church GM, Seidman CE, Seidman JG (2007) Polony multiplex analysis of gene expression (PMAGE) in a mouse model of hypertrophic cardiomyopathy. Science 316(5830):1481-4.
16. Church, GM (2006) Genomes for all. Scientific American Jan 2006.
15. Zhang, K, Martiny, AC, Reppas, NB, Barry, KW, Malek, J, Chisholm, SW, Church, GM (2006) Sequencing genomes from single cells via polymerase clones. Nature Biotech. Jun;24(6):680-6.
14. Fan JB, Chee, MS, Gunderson, KL (2006) Highly parallel genomic assays. Nature Reviews of Genetics 7:632-44. "Polonies are discrete clonal amplifications of a single DNA molecule, grown on a solid-phase surface. This approach greatly improves the signal-to-noise ratio. Polonies can be generated using several techniques that include solid-phase PCR in polyacrylamide gels, bridge PCR, rolling-circle amplification, BEAMing (beads, emulsions, amplification and magnetics)-based cloning on beads and massively parallel signature sequencing (MPSS) to generate clonal bead arrays."
13. Shendure, J, Porreca, GJ, Reppas, NB, Lin, X, McCutcheon, JP, Rosenbaum, AM, Wang, MD , Zhang, K, Mitra, RD, Church, GM (2005) Accurate Multiplex Polony Sequencing of an Evolved Bacterial Genome (Science 4-Aug) Science Supplement
12. Shendure J, Mitra R, Varma C, Church GM (2004) Advanced Sequencing Technologies: Methods and Goals. Nature Reviews of Genetics May;5(5):335-44.
11. Mikkilineni, V, Mitra, RD, Merritt, J, DiTonno, JR, Church, GM, Ogunnaike, B, and Edwards, JS. (2004) Digital Quantitative Measurements Of Gene Expression. Biotechnology and Bioengineering 86(2):117-24.
10. Dressman D, Yan H, Traverso G, Kinzler KW, and Vogelstein B. (2003) Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc Natl Acad Sci U S A. 2003 Jul 22;100(15):8817-22.
9. Butz J, Wickstrom E, and Edwards J. Characterization of mutations and loss of heterozygosity of p53 and K-ras2 in pancreatic cancer cell lines by immobilized polymerase chain reaction. BMC Biotechnol. 2003 Jul 23;3(1):11.
8. Constans, A (2003) Beyond Sanger: Toward the $1000 genome. The Scientist 17: 36.
7. Aach, J and Church, GM (2004) Mathematical models of diffusion-constrained polymerase chain reactions: basis of high-throughput nucleic acid assays and simple self-organizing systems. J. Theoret. Biol. May 7;228(1):31-46.
6. Buhler, J., Souvenir, R., Zhang, W, and Mitra, R. (2004) Design of a high-throughput assay for alternative splicing using polymerase colonies. PSB 2004.
5. Zhu,J, Shendure,J, Mitra, RD, and Church, GM (2003) Single Molecule Profiling of Alternative Pre-mRNA Splicing. Science. 2003 Aug 8;301(5634):836-8.
4. Mitra,RD, Shendure,J, Olejnik,J, Olejnik,EK, and Church,GM (2003) Fluorescent in situ Sequencing on Polymerase Colonies. Analyt. Biochem. 320:55-65.
3. Merritt J, DiTonno JR, Mitra RD, Church GM, and Edwards JS. (2003) Parallel competition analysis of Saccharomyces cerevisiae strains differing by a single base using polymerase colonies. . Nucleic Acids Res. Aug 1;31(15):e84.
2. Mitra, RD, Butty, V, Shendure, J, Williams, BR, Housman, DE, and Church, GM (2003) Digital Genotyping and Haplotyping with Polymerase Colonies. Proc Natl Acad Sci USA. May 13;100(10):5926-31.
1. Mitra, R. and Church, G.M. (1999) In situ localized amplification and contact replication of many individual DNA molecules. Nucleic Acids Res. 27(24):e34; pp.1-6.