





| Candidate UniGene cluster: | UniGene Cluster Hs.10882 |
| Description: | HMG-box containing protein 1 |
| Best-Of-UniGene (BOU) Sequence | NM_012257 |
| Genomic Coordiantes Displayed: | Bases 380000 to 520000 of contig Hs7_8071_24 |
| BOU Orientation Along Contig: | LEFT-TO-RIGHT with respect to contig |
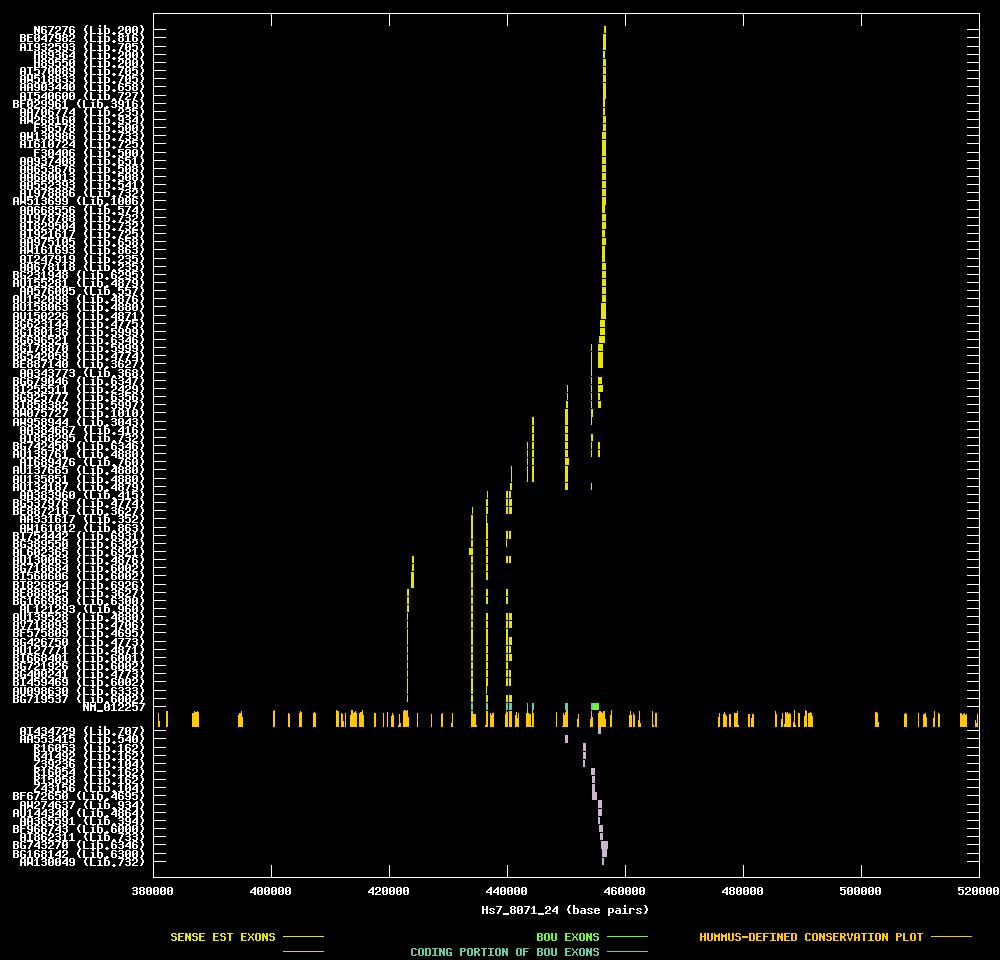
| Link to JPEG of genomic mapping | Hs.10882.jpeg |
| Best sense EST/protein match: | AI858295 matched ref|NP_036389.1| (NM_012257) HMG-box containing protein 1 [Homo sapiens] (E = e-108) |
| Best antisense EST/protein match: | No protein match with an e-value of less than 1e-10 |
ANTISENSE ESTs
| AI434729 | cDNA clone IMAGE:2132912 | lymph | 3' read | 0.6 kb |  | |
| AA553415 | cDNA clone IMAGE:1019504 | neural | 3' read | 4.2 kb |  | |
| R16053 | cDNA clone IMAGE:53072 | brain | 3' read | |||
| R41492 | cDNA clone IMAGE:29380 | brain | 3' read | 2.1 kb | ||
| Z39236 | cDNA clone c15a11 | brain | 3' read | |||
| R16054 | cDNA clone IMAGE:53072 | brain | 5' read |  | ||
| R15058 | cDNA clone IMAGE:29380 | brain | 5' read | 2.1 kb | ||
| Z43156 | cDNA clone c15a11 | brain | ||||
| BF672650 | cDNA clone IMAGE:4293130 | muscle (skeletal) | 5' read | |||
| AW274637 | cDNA clone IMAGE:2747296 | three pooled meningiomas | 3' read |  | ||
| AU144340 | cDNA clone HEMBA1001610 | whole embryo, mainly head | 3' read |  | ||
| AA365591 | cDNA clone ATCC:170339 | brain | 3' read |  | ||
| BF966743 | cDNA clone IMAGE:4375614 | hippocampus, cell line | 3' read | |||
| AI862311 | cDNA clone IMAGE:2265161 | uterus | 3' read | 0.4 kb | ||
| BG743270 | cDNA clone IMAGE:4778629 | skin | 5' read | |||
| BG168142 | cDNA clone IMAGE:4449535 | hypernephroma, cell line | 5' read | |||
| AW130049 | cDNA clone IMAGE:2619269 | uterus | 3' read |
| BG719537 | cDNA clone IMAGE:4822437 | testis, cell line | 5' read |  | ||
| AU098630 | cDNA clone COL05935 |  | ||||
| BI459469 | cDNA clone IMAGE:5266334 | testis, cell line | 5' read |  | ||
| BG400241 | cDNA clone IMAGE:4592743 | kidney | 5' read |  | ||
| BG721926 | cDNA clone IMAGE:4827901 | testis, cell line | 5' read |  | ||
| BI668401 | cDNA clone IMAGE:5311891 | hypothalamus, cell line | 5' read |  | ||
| AU127771 | cDNA clone NT2RP2002014 | 5' read |  | |||
| BG426750 | cDNA clone IMAGE:4606827 | kidney | 5' read |  | ||
| BF575809 | cDNA clone IMAGE:4289628 | muscle (skeletal) | 5' read |  | ||
| AV718093 | cDNA clone FHTAAAD11 | hypothalamus | 5' read |  |